 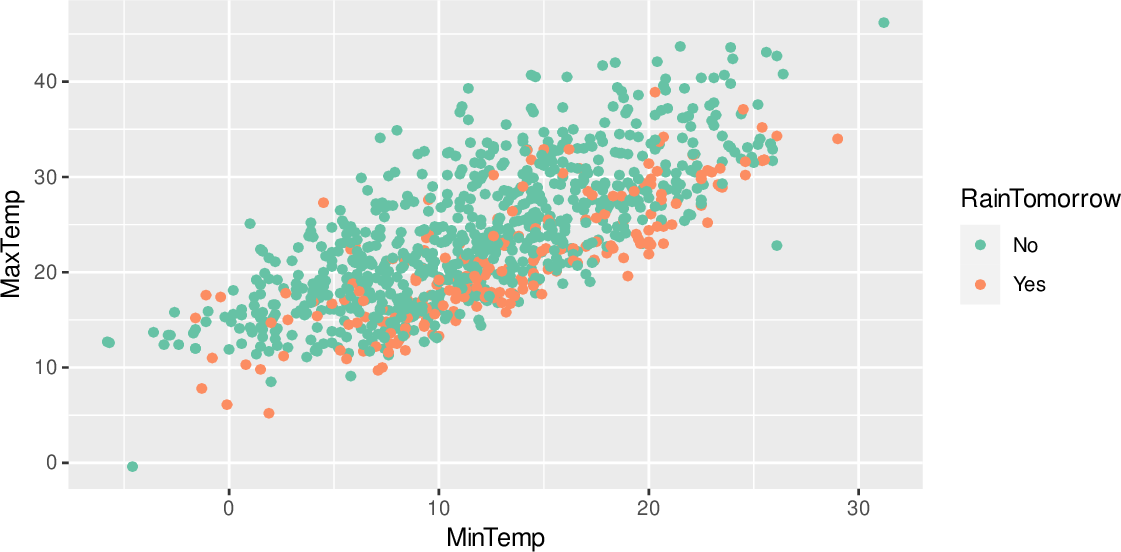
Normalize ): def _init_ ( self, vmin = None, vmax = None, midpoint = 0, clip = False ): self. score_genes ( adata, , score_name = "B_cell_score" ) # Palette normalization with centering and adapted dynamic range to correspond to # the distance of vmin and vmax from the cenetr # Adapted from class MidpointNormalize ( mcolors. # Centered non-symmetric palette # Make mock column for plotting, here we use B cell score sc. text ( x_loc, y_loc, gene, color = color_point, fontsize = 10 )) # Label selected genes on the plot _ = adjust_text ( texts, expand_points = ( 2, 2 ), arrowprops = dict ( arrowstyle = "->", color = "gray", lw = 1 ), ax = ax, ) set_title ( color ) # Labels # Select genes to be labeled texts = genes = for gene in genes : # Position of object to be marked x_loc = adata. scatter ( adata, x = x, y = y, color = color, show = False ) print ( "Axes:", ax ) # Move plot title from Axes to Legend ax. # In this example we want to show UMAPs of different cell type markers, # with markers of a single cell type in one row # and with a different number of markers per cell type (row) # Marker genes marker_genes = ): x = "means" y = "dispersions" color = "is_highly_variable" adata. with show=False) we obtain either an individual Axes object (if this is the only Axes object on the Figure) or a list of Axes (if multiple Axes were created). When accessing Axes from Figure the returned object is a list and we need to select the relevant Axes to modify them. However, if we want to obtain the colorbar axes object we need to use return_fig=True rather than show=False. For every plotted category one Axes object will be created and for every continuous category two Axes objects: the UMAP plot and colorbar on the side. If we want to customize Axes after the scanpy plotting function was called we need to set show=False to ensure that the plot will be rendered only after we made all adjustments.įor example, from embedding plots (such as umap) we can obtain either axes (by setting show=False) or the whole figure (by setting return_fig=True) that stores axes in figure.axes. The show parameter also regulates when the plot is rendered. Scanpy plotting functions can return Figure or the plot object (by setting return_fig=True) or Axes (by setting show=False). A note on Figures and Axes used in Scanpy plots # continous Colorbar or discrete Legend), etc. There are also other differences, such as which types of legends are used (i.e. 
Certain functions plot on individual Axes objects while others use the whole Figure, combining multiple Axes to display different parts of a single plot. Please note that some tutorial parts are specific for individual scanpy ploting functions, as they create plots in different ways. Some scanpy functions can also take as an input predefined Axes, as shown below. Matplotlib plots are drawn in Figure objects which in turn contain one or multiple Axes objects. Scanpy plots are based on matplotlib objects, which we can obtain from scanpy functions and subsequently customize. This section provides general information on how to customize plots. pbmc68k_reduced () Adapting matplotlib based plots # ggplot ( mtcars2, aes ( wt, mpg ) ) + geom_point (na.Adata = sc. P Warning: Removed 4 rows containing missing values or values outside the scale #> range (`geom_point()`). That define both data and aesthetics and shouldn't inherit behaviour from If FALSE, overrides the default aesthetics, 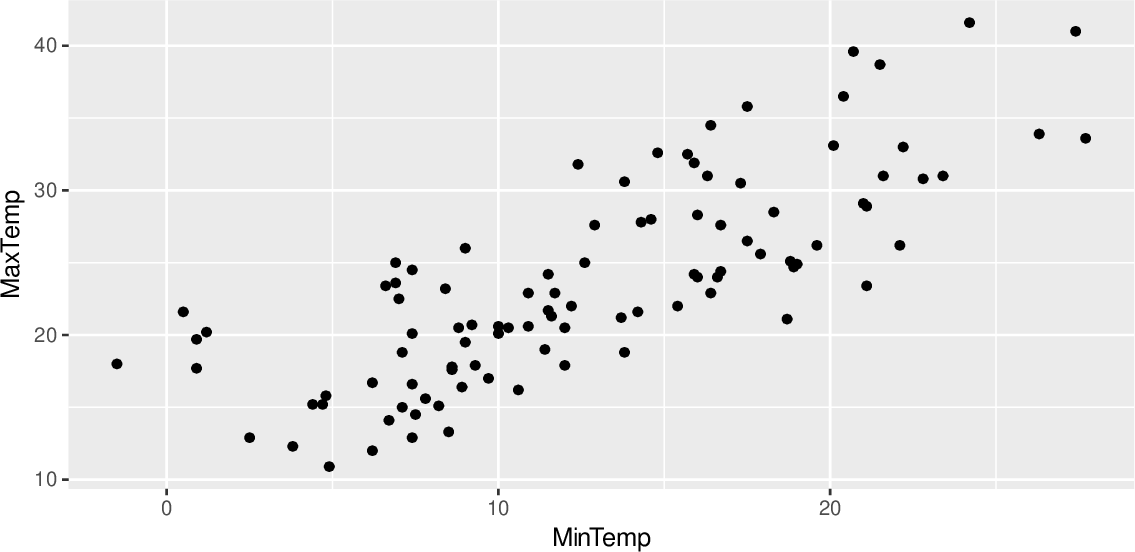
It can also be a named logical vector to finely select the aesthetics to NA, the default, includes if any aesthetics are mapped.įALSE never includes, and TRUE always includes. Should this layer be included in the legends? If TRUE, missing values are silently removed. If FALSE, the default, missing values are removed withĪ warning. Often aesthetics, used to set an aesthetic to a fixed value, likeĬolour = "red" or size = 3. "jitter" to use position_jitter), or the result of a call to a Position adjustment, either as a string naming the adjustment Layer, either as a ggproto Geom subclass or as a string naming the 
The statistical transformation to use on the data for this A function can be createdįrom a formula (e.g. Seeįortify() for which variables will be created.Ī function will be called with a single argument, All objects will be fortified to produce a data frame. If NULL, the default, the data is inherited from the plotĭata as specified in the call to ggplot().Ī ame, or other object, will override the plotĭata. You must supply mapping if there is no plot Inherit.aes = TRUE (the default), it is combined with the default mappingĪt the top level of the plot. Set of aesthetic mappings created by aes(). 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
